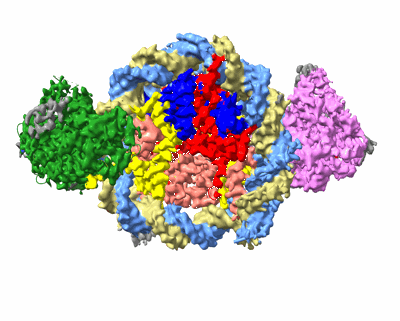
Structure of DDM1-nucleosome complex in the apo state
Liu Y, Zhang Z, Hu H, Chen W, Zhang F, Wang Q, Wang C, Yan K, Du J
Nat Plants (2024) 10 pp. 374-380 [ Pubmed: 38413824 DOI: doi:10.1038/s41477-024-01640-z ]
- Atp-dependent dna helicase ddm1 (86 kDa, Protein from Arabidopsis thaliana)
- Histone h2b.6 (16 kDa, Protein from Arabidopsis thaliana)
- Dna (antisense strand) (51 kDa, DNA from synthetic construct)
- Histone h3.1 (15 kDa, Protein from Arabidopsis thaliana)
- Histone h4 (11 kDa, Protein from Arabidopsis thaliana)
- Dna (sense strand) (51 kDa, DNA from synthetic construct)
- Ddm1-nucleosome complex in the apo state (290 kDa, Complex from Arabidopsis thaliana)
- Histone h2a.6 (13 kDa, Protein from Arabidopsis thaliana)
Structure of DDM1-nucleosome complex in ADP state
Resolution: 3.6 Å
EM Method: Single-particle
Fitted PDBs: 8wh8
Q-score: 0.403
Liu Y, Zhang Z, Hu H, Chen W, Zhang F, Wang Q, Wang C, Yan K, Du J
Nat Plants (2024) 10 pp. 374-380 [ Pubmed: 38413824 DOI: doi:10.1038/s41477-024-01640-z ]
- Atp-dependent dna helicase ddm1 (86 kDa, Protein from Arabidopsis thaliana)
- Histone h2b.6 (16 kDa, Protein from Arabidopsis thaliana)
- Dna (sense strand) (45 kDa, DNA from synthetic construct)
- Dna (antisense strand) (45 kDa, DNA from synthetic construct)
- Histone h3.1 (15 kDa, Protein from Arabidopsis thaliana)
- Histone h4 (11 kDa, Protein from Arabidopsis thaliana)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Ddm1-nucleosome complex in adp state (290 kDa, Complex from Arabidopsis thaliana)
- Histone h2a.6 (13 kDa, Protein from Arabidopsis thaliana)
Structure of DDM1-nucleosome complex in ADP-BeFx state
Resolution: 3.31 Å
EM Method: Single-particle
Fitted PDBs: 8wh9
Q-score: 0.336
Liu Y, Zhang Z, Hu H, Chen W, Zhang F, Wang Q, Wang C, Yan K, Du J
Nat Plants (2024) 10 pp. 374-380 [ Pubmed: 38413824 DOI: doi:10.1038/s41477-024-01640-z ]
- Atp-dependent dna helicase ddm1 (86 kDa, Protein from Arabidopsis thaliana)
- Histone h2b.6 (16 kDa, Protein from Arabidopsis thaliana)
- Dna (sense strand) (45 kDa, DNA from synthetic construct)
- Dna (antisense strand) (45 kDa, DNA from synthetic construct)
- Histone h3.1 (15 kDa, Protein from Arabidopsis thaliana)
- Histone h4 (11 kDa, Protein from Arabidopsis thaliana)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Beryllium trifluoride ion (66 Da, Ligand)
- Ddm1-nucleosome complex in adp-befx state (290 kDa, Complex from Arabidopsis thaliana)
- Histone h2a.6 (13 kDa, Protein from Arabidopsis thaliana)
Structure of DDM1-nucleosome complex in the ADP-BeFx state with DDM1 bound to SHL2 and SHL-2
Resolution: 4.05 Å
EM Method: Single-particle
Fitted PDBs: 8wha
Q-score: 0.306
Liu Y, Zhang Z, Hu H, Chen W, Zhang F, Wang Q, Wang C, Yan K, Du J
Nat Plants (2024) 10 pp. 374-380 [ Pubmed: 38413824 DOI: doi:10.1038/s41477-024-01640-z ]
- Ddm1-nucleosome complex in the adp-befx state with ddm1 bound to shl2 and shl-2 (376 kDa, Complex from Arabidopsis thaliana)
- Histone h2b.6 (16 kDa, Protein from Arabidopsis thaliana)
- Dna (sense strand) (45 kDa, DNA from synthetic construct)
- Dna (antisense strand) (45 kDa, DNA from synthetic construct)
- Atp-dependent dna helicase ddm1 (86 kDa, Protein from Arabidopsis thaliana)
- Histone h3.1 (15 kDa, Protein from Arabidopsis thaliana)
- Histone h4 (11 kDa, Protein from Arabidopsis thaliana)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Beryllium trifluoride ion (66 Da, Ligand)
- Histone h2a.6 (13 kDa, Protein from Arabidopsis thaliana)
Structure of nucleosome core particle of Arabidopsis thaliana
Resolution: 3.17 Å
EM Method: Single-particle
Fitted PDBs: 8whb
Q-score: 0.5
Liu Y, Zhang Z, Hu H, Chen W, Zhang F, Wang Q, Wang C, Yan K, Du J
Nat Plants (2024) 10 pp. 374-380 [ Pubmed: 38413824 DOI: doi:10.1038/s41477-024-01640-z ]
- Histone h2b.6 (16 kDa, Protein from Arabidopsis thaliana)
- Structure of the nucleosome core particle of arabidopsis thaliana (200 kDa, Complex from Arabidopsis thaliana)
- Dna (antisense strand) (45 kDa, DNA from synthetic construct)
- Histone h3.1 (15 kDa, Protein from Arabidopsis thaliana)
- Histone h4 (11 kDa, Protein from Arabidopsis thaliana)
- Dna (sense strand) (45 kDa, DNA from Arabidopsis thaliana)
- Histone h2a.6 (13 kDa, Protein from Arabidopsis thaliana)
Cryo-EM structure of Oryza sativa HKT2;1 at 2.5 angstrom
Resolution: 2.53 Å
EM Method: Single-particle
Fitted PDBs: 8k66
Q-score: 0.66
Wang X, Shen X, Qu Y, Zhang H, Wang C, Yang F, Shen H
Nat Plants (2024) 10 pp. 633-644 [ DOI: doi:10.1038/s41477-024-01665-4 Pubmed: 38570642 ]
- 1,2-diacyl-sn-glycero-3-phosphocholine (790 Da, Ligand)
- Cholesterol hemisuccinate (486 Da, Ligand)
- Sodium ion (22 Da, Ligand)
- Water (18 Da, Ligand)
- Cation transporter hkt2;1 (60 kDa, Protein from Oryza sativa subsp. japonica)
- Phosphatidylinositol (887 Da, Ligand)
- Oshkt2;1 dimer (Complex from Oryza sativa subsp. japonica)
- Phosphatidylethanolamine (734 Da, Ligand)
- Cholesterol (386 Da, Ligand)
Cryo-EM structure of Oryza sativa HKT2;2/1 at 2.3 angstrom
Resolution: 2.33 Å
EM Method: Single-particle
Fitted PDBs: 8k69
Q-score: 0.692
Wang X, Shen X, Qu Y, Zhang H, Wang C, Yang F, Shen H
Nat Plants (2024) 10 pp. 633-644 [ DOI: doi:10.1038/s41477-024-01665-4 Pubmed: 38570642 ]
- Oshkt2;2/1 dimer (Complex from Oryza sativa subsp. indica)
- 1,2-diacyl-sn-glycero-3-phosphocholine (790 Da, Ligand)
- Cholesterol hemisuccinate (486 Da, Ligand)
- Sodium ion (22 Da, Ligand)
- Water (18 Da, Ligand)
- Phosphatidylinositol (887 Da, Ligand)
- Cation transporter hkt2;2 (60 kDa, Protein from Oryza sativa subsp. indica)
- Phosphatidylethanolamine (734 Da, Ligand)
- Cholesterol (386 Da, Ligand)
Apo state of Arabidopsis AZG1 at pH 7.4
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 8irl
Q-score: 0.604
Xu L, Jia W, Tao X, Ye F, Zhang Y, Ding ZJ, Zheng SJ, Qiao S, Su N, Zhang Y, Wu S, Guo J
Nat Plants (2024) 10 pp. 180-191 [ Pubmed: 38172575 DOI: doi:10.1038/s41477-023-01590-y ]
- Adenine/guanine permease azg1 (65 kDa, Protein from Arabidopsis thaliana)
- Azg1 dimer (131 kDa, Complex from Arabidopsis thaliana)
Endogenous substrate adenine bound state of Arabidopsis AZG1 at pH 5.5
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8irm
Q-score: 0.601
Xu L, Jia W, Tao X, Ye F, Zhang Y, Ding ZJ, Zheng SJ, Qiao S, Su N, Zhang Y, Wu S, Guo J
Nat Plants (2024) 10 pp. 180-191 [ Pubmed: 38172575 DOI: doi:10.1038/s41477-023-01590-y ]
- Water (18 Da, Ligand)
- Adenine/guanine permease azg1 (65 kDa, Protein from Arabidopsis thaliana)
- Complex of azg1 and adenine (131 kDa, Complex from Arabidopsis thaliana)
- Adenine (135 Da, Ligand)
6-BAP bound state of Arabidopsis AZG1
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 8irn
Q-score: 0.595
Xu L, Jia W, Tao X, Ye F, Zhang Y, Ding ZJ, Zheng SJ, Qiao S, Su N, Zhang Y, Wu S, Guo J
Nat Plants (2024) 10 pp. 180-191 [ Pubmed: 38172575 DOI: doi:10.1038/s41477-023-01590-y ]
- Water (18 Da, Ligand)
- Complex of azg1 and 6-bap (131 kDa, Complex from Arabidopsis thaliana)
- Adenine/guanine permease azg1 (65 kDa, Protein from Arabidopsis thaliana)
- N-benzyl-9h-purin-6-amine (225 Da, Ligand)